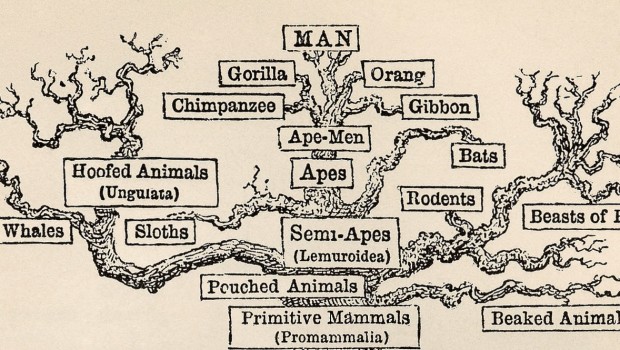
Episode 26: The Tree of Mammals
Mammals are an incredibly diverse and highly successful group of animals. They include some of the tallest, heaviest and fastest animals around today, as well as our own species. For over 100 years, biologists have attempted to build mammal evolutionary trees using anatomical data. This work has provided the basis for our understanding of mammal relationships. Within the last 30 years, new technologies have enabled scientists to cheaply sequence molecular data (e.g. DNA and amino acid sequences) from thousands of mammal species. Interestingly, molecular trees reveal close relationships between some very different looking mammals. To guide us through this mammal renaissance, we are joined by Dr Robert Asher from the Department of Zoology at the University of Cambridge, UK.
Podcast: Download (Duration: 43:56 — 40.2MB)
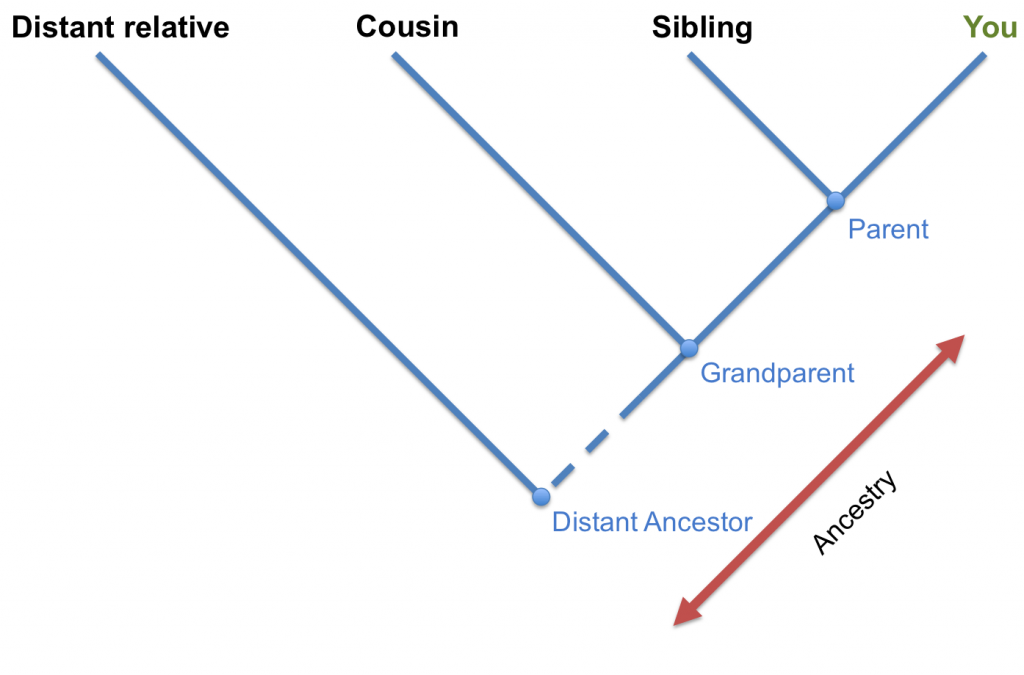
Building the Tree of Life is just like building a family tree. Close relatives share recent common ancestors (for example siblings share parents, cousins share grandparents), where as distant relatives share less recent common ancestors (for example second cousins share great-grandparents, third cousins share great-great-grandparents). In the same way, Humans and Chimpanzees share a recent ape ancestor, where as humans and Gibbons share a more distant ape ancestor.
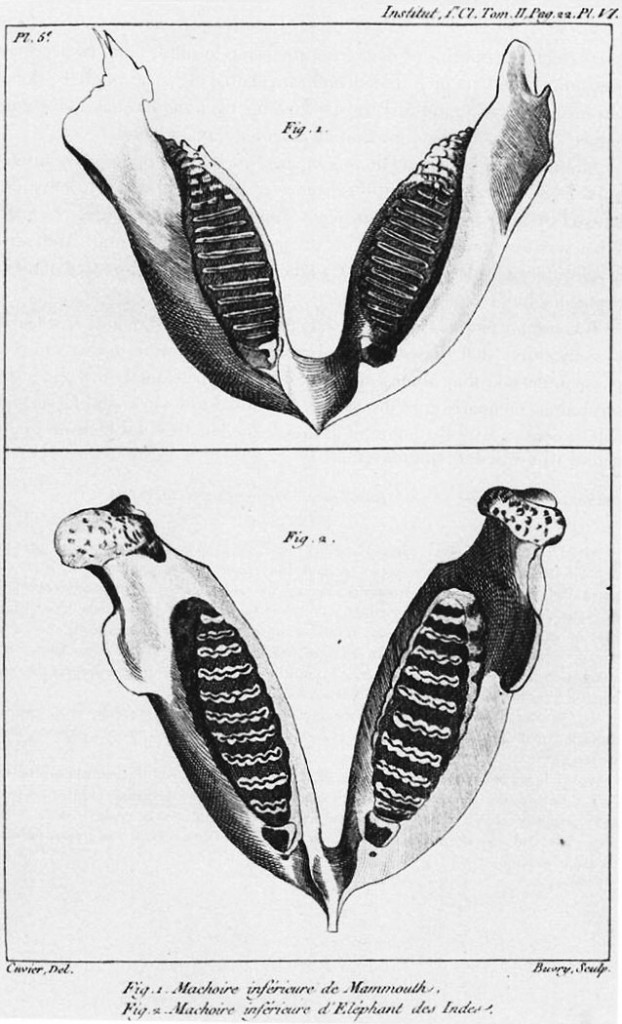
Anatomical data can be used to classify both living and fossil organisms. One of the first people who attempted to classify fossils with living animals was the French zoologist Georges Cuvier. In the figure above, Cuvier compares the jaw of an Indian Elephant with that of an extinct mammoth. From Cuvier 1969-1832 PD-US.
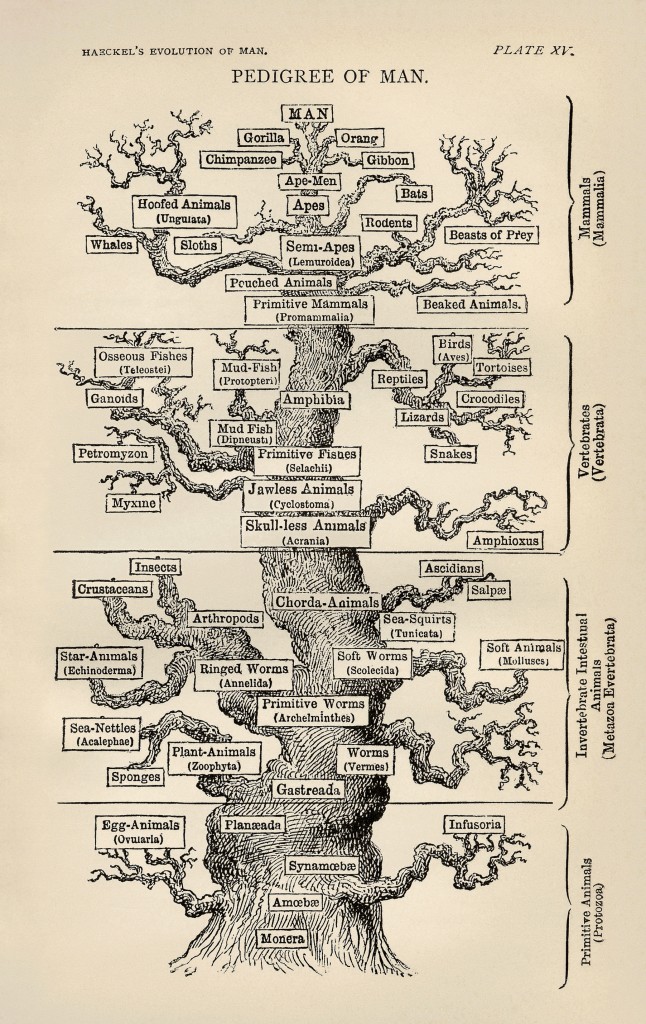
Morphological and anatomical data can be used to build evolutionary trees. Some of the first were drawn by the German biologist Ernst Haeckel. From Haeckel 1879 PD-US.
Genetic sequences and amino acids are another source of data from which you can build evolutionary trees. This kind of data wasn’t available to the evolutionary biologists of the 19th century such as Darwin and Haeckel. In contrast, websites like GenBank store genetic information for hundreds of thousands of organisms. This data is freely available which means that, if you wanted to, you could download the entire genome of an African Elephant in a couple of clicks.
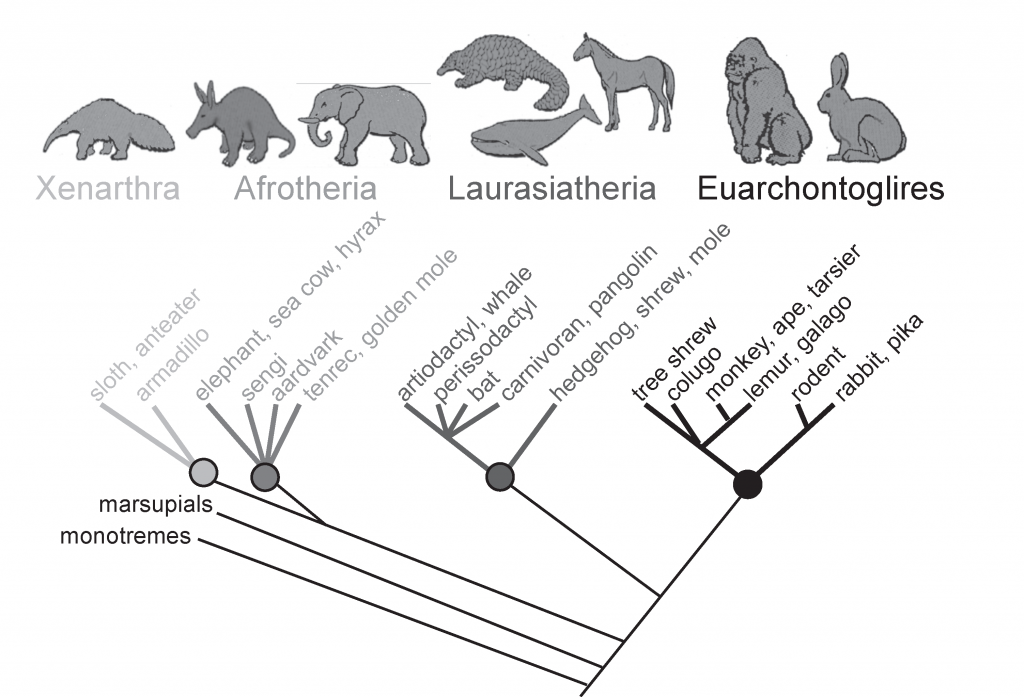
The inclusion of molecular data from DNA and/or protein sequences has enhanced our understanding of the mammal tree. However, the overall shape of the tree of mammals hasn’t drastically changed since the first trees were drawn in the 19th Century. For example, molecular data still supports a grouping of hoofed mammals. On the other hand, molecular data has produced some unexpected branches on the mammal tree. One surprising result is that molecular data suggests a number of African mammals with little morphological resemblance are, in fact, closely related. This group is called ‘Afrotheria’. From Asher 2012 Evolution & Belief.
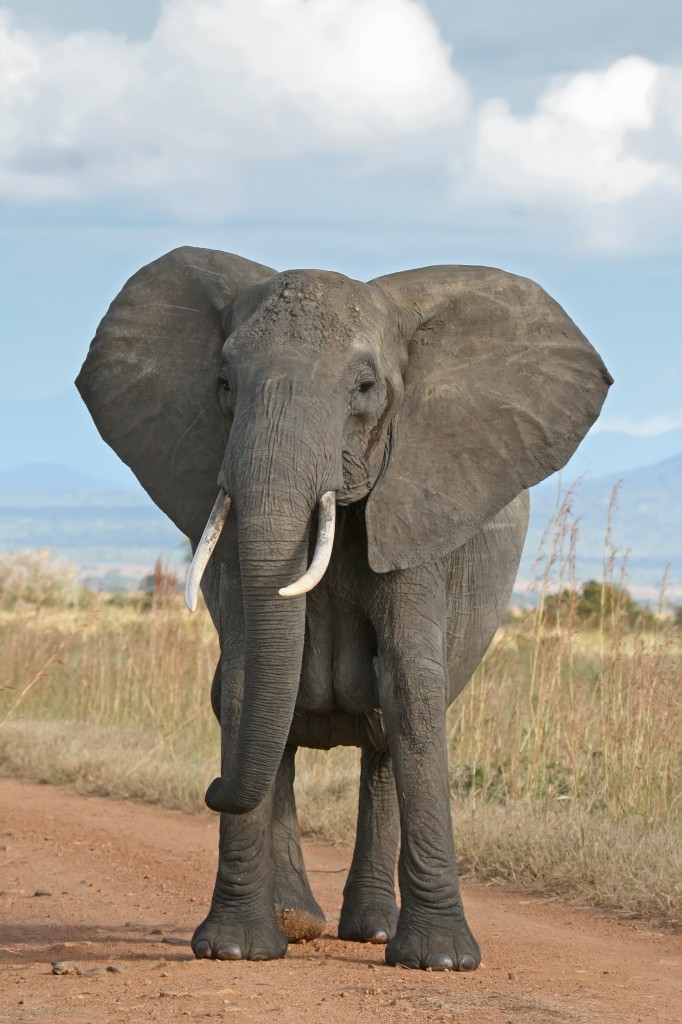

Afrotheria includes the largest living land mammals; the elephants. From Wikimedia Commons, GNU Free Documentation License.
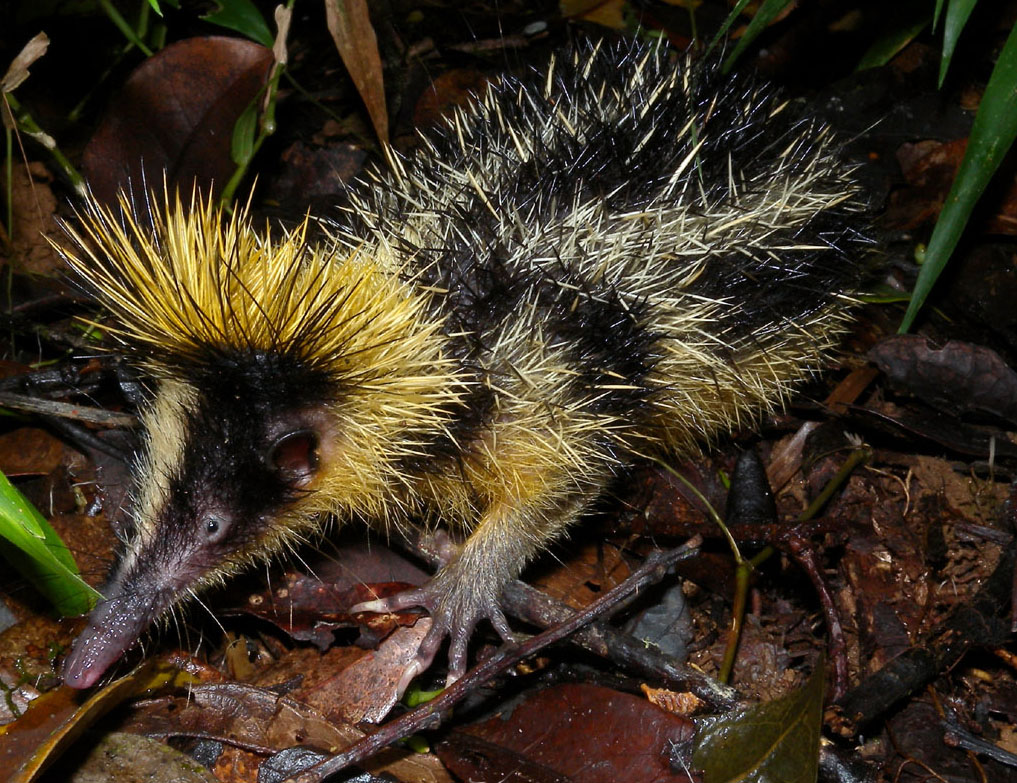

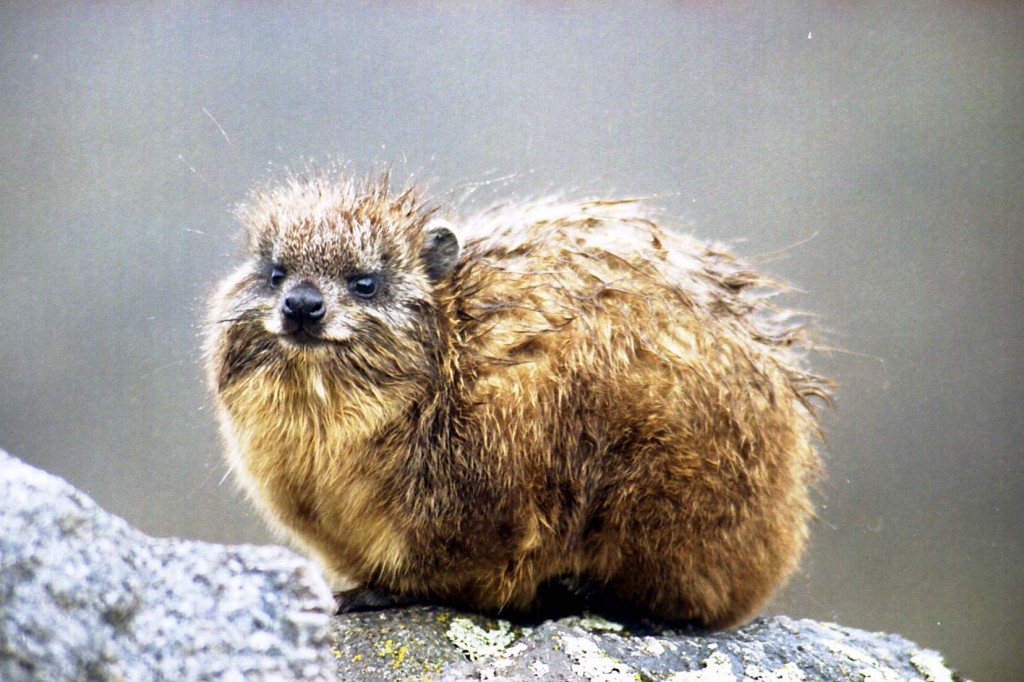
This Lowland Streaked Tenrec is native to Madagascar. While it might resemble hedgehogs or shrews, it is an afrotherian, so it is more closely related to elephants. Photo by Frank Vassen, licensed under CC.

Similarly this Golden Mole is an afrotherian. While it might resemble true moles, it is actually closely related to tenrecs. From Wikimedia Commons, GNU Free Documentation License.
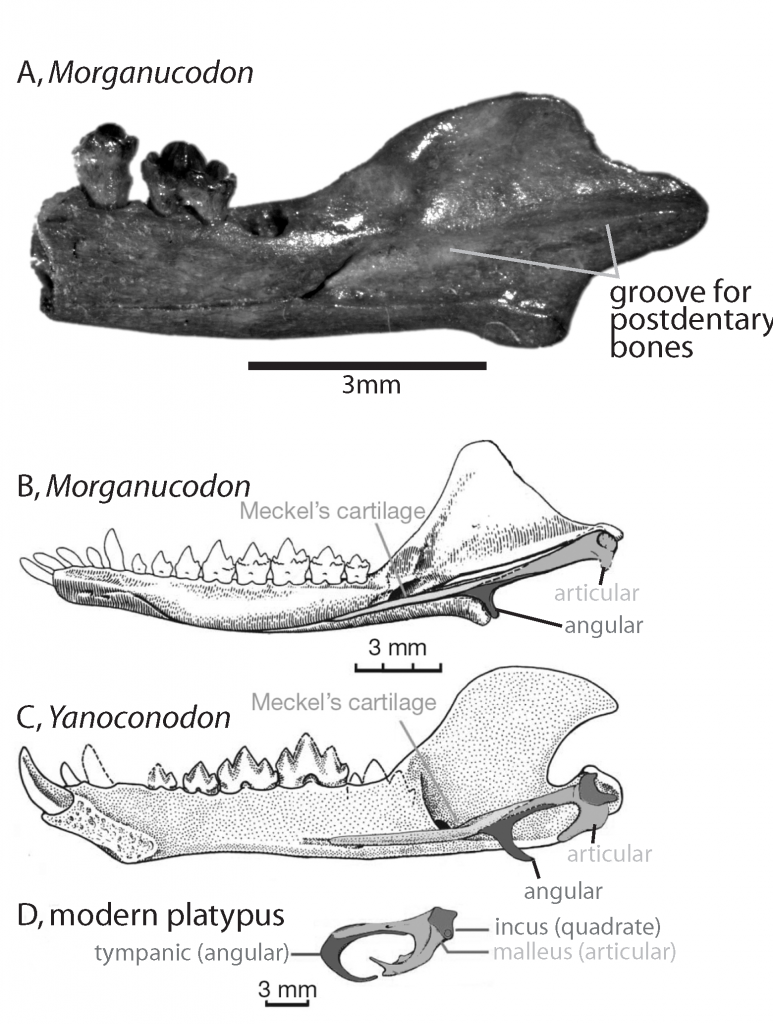
Mammals have a good fossil record. This is particularly true if we consider the interval during which the mammal body-plan evolved. Morganucodon and Yanoconodon are thought to be closely related to living mammals because they share a number of anatomical characteristics such as discrete bands of differently shaped teeth. Morganucodon is thought to be more closely related to mammals than is Yanoconodon because it has a mammal-like jaw articulation. From Asher 2012 Evolution & Belief.
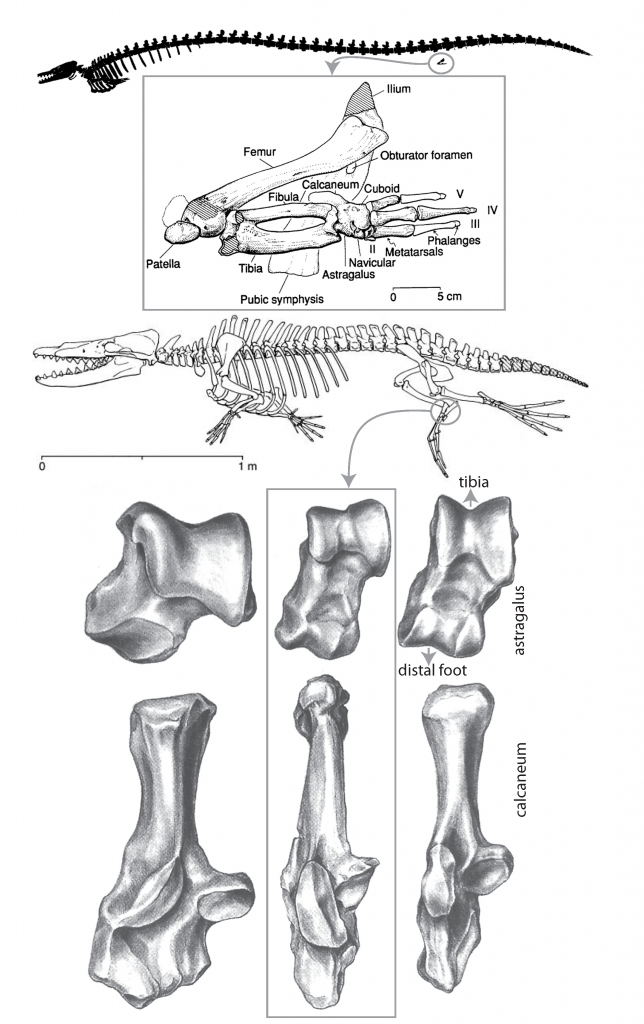
The evolution of whales is another area where the fossil record of mammals excels. The early whale Basilosaurus shows greatly reduced hindlimbs that clearly would not have functioned as a weight bearing appendages. Intriguingly the hindlimb of Basilosaurus shows that digits III and IV are the largest. This is the same digit pattern that we see in living even-toed ungulates (even toed hoofed mammals) and betrays the close evolutionary relationship between whales and hoofed mammals. From Asher 2012 Evolution & Belief.
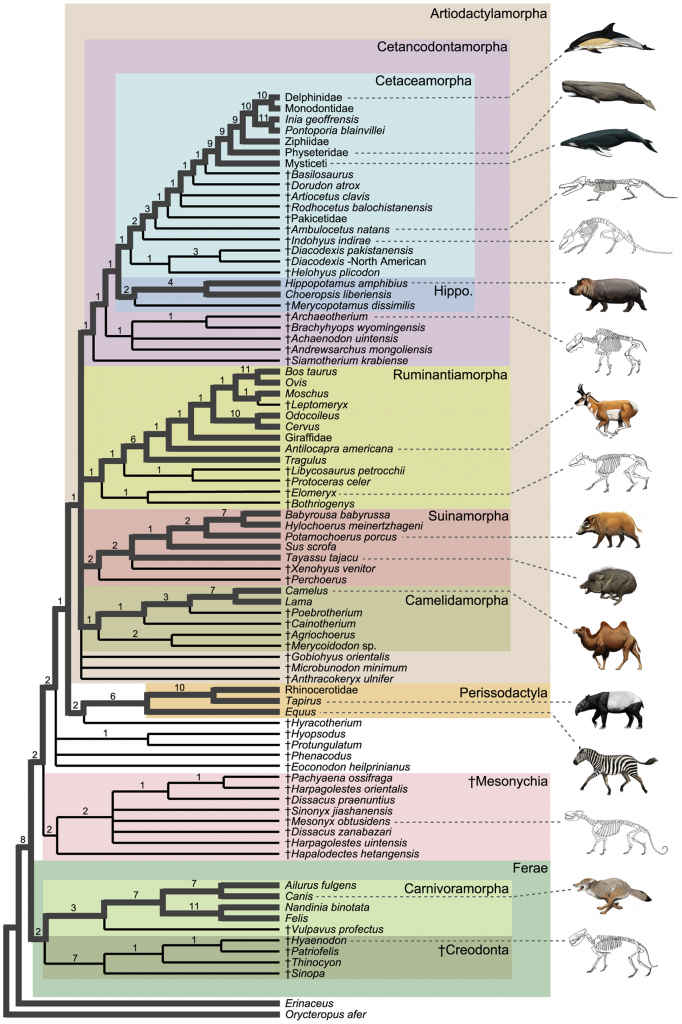
Molecular data reveals that hippos are the closest living relatives of whales. However, the fossil record bridges the gap between hippos and whales with a number of extinct lineages. These extinct whale-relatives record the stepwise evolution of whale-like traits. From Spaulding et al. 2009, licensed under CC.